Recently, the Vegetable Functional Genomics Innovation Team at the Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, published a research paper online in the internationally renowned academic journal Microbiome (IF-5y = 16.6), titled "Host Genetic Regulation of Xylem-Resident Pseudomonas Enhances Cucumber Growth." This study systematically dissects the genetic basis of the cucumber xylem microbial composition and reveals a novel mechanism by which the host gene CsXPR1 promotes cucumber growth through the specific recruitment of beneficial Pseudomonas. It emphatically elucidates that this growth-promoting effect exhibits significant host genotype dependency.

Endophytic microorganisms play a key role in plant growth and stress resistance. However, the genetic basis of how plants precisely select and recruit these beneficial microbes, especially within the xylem responsible for long-distance transport of water and nutrients, has remained unclear. This study conducted a genome-wide association study (GWAS) of the xylem microbiome on 109 cucumber core accessions with extensive genetic diversity. It was discovered that the cucumber xylem harbors a highly conserved core microbial community, dominated by the genus Pseudomonas.
The research team further pinpointed a key host genetic locus, CsXPR1, through GWAS. This gene encodes a TPR (Tetratricopeptide Repeat) protein that precisely regulates the abundance of Pseudomonas fulva strain 220, a core beneficial bacterium in the xylem. More importantly, the study revealed that this microbial growth-promoting effect is significantly "genotype-dependent." Mechanistic studies showed that after colonizing the xylem, P. fulva strain 220 significantly increases nitrogen levels in the xylem sap by synthesizing the key metabolite 4-methyleneglutamine, thereby promoting plant growth. Inoculation experiments confirmed that this strain only exhibits a significant growth-promoting effect in the cucumber haplotype with high CsXPR1 expression (Hap2), leading to notable increases in plant height, stem diameter, leaf area, and biomass. Conversely, in the haplotype with low CsXPR1 expression (Hap1), where the host's ability to recruit this strain is relatively weak, the growth-promoting effect was not significant. This finding elucidates the interaction mechanism of "precise recruitment and co-evolution" between plants and their core microbiome (Figure 1).
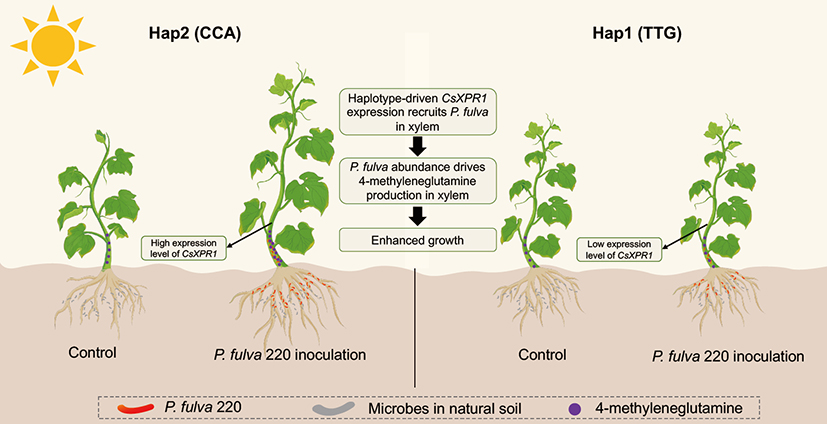
Figure 1: Genotype-dependent growth promotion mechanism by xylem-resident Pseudomonas
This research achievement not only fills a gap in the study of how host genetic variation reshapes the xylem endophytic community but also constructs a complete "host gene-beneficial microbe-metabolite-plant phenotype" functional pathway. Importantly, it provides crucial genetic resources and a theoretical foundation for the precise utilization of the plant microbiome through genetic improvement and for breeding new, high-yielding, and efficient crop varieties. The precise interaction model of "specific genotype pairing with specific bacteria" revealed by this study outlines a blueprint for the future evolution of agriculture from traditional single-factor breeding towards the synergistic co-evolution of "germplasm + microbiome." This precise customization model based on the Host-Genotype-Microbiome implies that in the future, we can tailor functional microbial consortia according to the genetic background of crops, building a new type of intelligent agricultural ecosystem. Through the precise customization of germplasm resources and the comprehensive utilization of highly beneficial microbial communities, agricultural production will embark on a more green, efficient, and sustainable development track.
This research was conducted with the Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, as the primary and corresponding author institution. Dr. Qin Yuxuan, former master's student Zhu Xueying, current Ph.D. candidate Zheng Yingying, and former Ph.D. graduate Wang Kun from the institute are co-first authors of the paper. Professor Yang Xueyong and Associate Professor Qin Yuxuan from the Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, and Professor Wei Hailei from the Institute of Agricultural Resources and Regional Planning, Chinese Academy of Agricultural Sciences, are co-corresponding authors. Professor Yang Jinliang from the University of Nebraska-Lincoln, USA, participated in the research. Professor Ai Chao from the Institute of Agricultural Resources and Regional Planning, Chinese Academy of Agricultural Sciences, provided valuable suggestions for this study. This research was supported by the National Key R&D Program of China, the National Natural Science Foundation of China, and the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences.
Original Link: https://link.springer.com/article/10.1186/s40168-025-02308-2